A not complete test analysis of the behavior of GATTS is also available.
If you want to see how things are done in more detail, feel free to take a look on the source files.
 Information
InformationGATSS is a Genetic Algorithm based solver of the Traveling Salesman problem written in GNU C++ linked to a user interface in HTML via CGI-script. It does nothing revolutional to the science of GA, it is just a standard algorithm enclosed in a user friendly interface. Maybe it can be used as a tool in teaching the concepts of Genetic Algorithms.
This program and its documentation is the result of an exercise in writing some kind of application related to the science of Artificial Life. The exercise is an examination part of the course "Artificial Intelligence II" given by the Department of Computing Science at Umeĺ University, Sweden.
The Story
When I first heard about Genetic Algorithms, I was fascinated by the fact that for once, the nature was the role model. That something as formal and abstract as Computer Science could learn from nature.
I searched the web for information on GA:s and found several interesting applications, general GA problem solvers, as well as GA software engineering environments.
One of the applications i found was 'DOUGAL' written by Brett S. Parker in 1993. It is a GA demonstration program written in Pascal for MS-DOS solving the Traveling Salesman problem (hereafter called TSP). I thought that the TSP was a good example in showing how GA works. It was a well known graph-theoretical problem and the results could, obviously, be nice visualised.
GATSS is very similar to DOUGAL. The differences are mainly that GATSS is implemented in C++ and has a HTML user interface.
So, thank you Brett, I hope you don't mind!.
When I had decided what kind of problem to solve (TSP) and in what language (C++), I searched for software engineering tools to speed up development. My first stop was not beautiful. I won't mention the name of the system here but it was ugly. Bad written code and bad written documentation.
 Later on I ran into a C++/perl GA software engineering environment called 'GAGS' written by professor J.J. Merelo, GeNeura Team at the Department of Electronics and Computer Technology, University of Granada, Spain. I had difficulties in compiling GAGS for two reasons, one: I got the latest beta-version (with link bugs) at the time, two: The system had never been compiled at the UNIX-platform I used. Mr. Merelo was very helpful, and after some email-correspondance, the system was up and running.
Later on I ran into a C++/perl GA software engineering environment called 'GAGS' written by professor J.J. Merelo, GeNeura Team at the Department of Electronics and Computer Technology, University of Granada, Spain. I had difficulties in compiling GAGS for two reasons, one: I got the latest beta-version (with link bugs) at the time, two: The system had never been compiled at the UNIX-platform I used. Mr. Merelo was very helpful, and after some email-correspondance, the system was up and running.
Since I'm no C++-guru, and Merelo really uses the language at it's full extent, together with the fact that I'm new to the field of GA, I had difficulties in understanding the code. I finally decided to create the code I needed from scratch instead. No shadow on the GAGS-system though. It is a well written general C++-library combined with perl scripts, with program examples and a nice user interface. The documentation is at the time a little poor (at least the programmers manual) but Merelo says he's working on it.
Thank you J.J. for the help in compiling GAGS! Even if I couldn't use the system itself, it has inspired my own code writing.
When the actual coding was finished, phase two of this project was to be done: Analysis of the different parameters in the GA, i.e. what effect they have on the evolution process. This is greatly simplified with a good user interface. To execute the original program, the command line would look like this:
> travel 100 10 1.0 0 0.5 1 0 5 0resulting in
Using seed: 799759313 Data: [0] average: 66.8872 best: 52.477 [1] average: 58.9228 best: 48.2341 [2] average: 56.191 best: 48.2341 [3] average: 54.6087 best: 48.2341 [4] average: 53.034 best: 48.2341 [5] average: 51.8041 best: 48.2341 [6] average: 50.38 best: 48.2341 [7] average: 49.4576 best: 48.2341 [8] average: 48.5745 best: 48.2341 [9] average: 48.4072 best: 48.2341 End of Data Best solution: [0]: [0]: Umeĺ (0,0) [1]: [1]: Stockholm (-1,-5) [2]: [4]: London (-8,-12) [3]: [5]: Berlin (-2,-11) [4]: [6]: Prag (-1,-14) [5]: [3]: Moskva (9,-9) [6]: [2]: Helsingfors (3,-4) [7]: [7]: Umeĺ (0,0) Distance: 48.2341
The difficulties in keeping track of which of the parameters that was corresponding to which part of the GA made me dizzy. I decided to create a better user interface using HTML.
Here, WWW-guru Daniel Silén, helped me out. He wrote the CGI-script that links the user interface to the program. Thank you Daniel!
The most important parts of a GA is crossover, mutation, population deletion, and chromosome fitness evaluation.
Mutation
The purpose of mutation is to reintroduce divergence into a converging population. This is a tool in avoiding local maxima, which is a common problem in stochastic algorithms. The mutation can be thought of as an error in the reproduction process. The thing is that this error is very important since it is the only possible way to create truly new chromosomes (crossover uses old gene material).
Mutation is applied in combination with crossover.
Population Deletion
This GA part defines how new chromosomes will be put into the existing population.
In general this can be done in at least two ways. You can either put the new chromosomes into an empty population and, when finished mating, make the new population the current population, or you can update the current population continuosly during the mating process, allowing new chromosomes act like parents to children in their own generation!
The former deletion technique is called 'Delete All' and the latter 'Steady Delete'. Both these techniques are provided in GATSS.
Fitness Evaluation
The evaluation algorithm used is crucial to how well the GA manages to find a good solution. If, like in Traveling Salesman, the structure of a good solution is well known (the sum of the distance between nodes shold be as small as possible), it is quite easy to construct a suitable algorithm. There are, however problems that are more tricky to give a good fitness evaluation algorithm.
The fitness evaluation of chromosomes is very problem dependent.
Data Structures
The data structures used in GA are usually rough simplifications of their natural prototype. Genes, chromosomes and populations are designed hierarchically somewhat like this:

More information about Genetic Algorithms is available at The Genetic Algorithms Archive or in the comp.ai.genetic FAQ's.
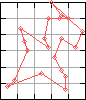 The Traveling Salesman problem (TS hereafter) is a classical graph-theoretical problem. It involves a traveling salesman who has to visit each of the cities in a given network before returning to his starting point, at which time his trip is complete. The algorithmic problem asks for the cheapest route, namely for a closed tour that passes through each of the nodes in the graph and whose total cost (i.e. the sum of the distances labeling the edges) is minimal.
The Traveling Salesman problem (TS hereafter) is a classical graph-theoretical problem. It involves a traveling salesman who has to visit each of the cities in a given network before returning to his starting point, at which time his trip is complete. The algorithmic problem asks for the cheapest route, namely for a closed tour that passes through each of the nodes in the graph and whose total cost (i.e. the sum of the distances labeling the edges) is minimal.
Its variants arise in the design of telephone networks and integrating circuits, in planning construction lines, and in the programming of industrial robots, to mention but a few applications. In all of these, the ability to find inexpensive tours of the graphs in question can be quite crucial.
TS is a NPC (Nondeterministic Polynomial-time Complete) problem which in short means that it is unsolvable with a standard linear algorithm if the problems complexity is large enough. TS can be solved with a very fast computer when the number of nodes are up to, say 50. If the graph is larger, a non-linear algorithm is needed. GA is one alternative.
Crossover
The crossover types used in GATSS are called Fixed Position and Fixed Position Sequential:
Fixed Position
The genes of parent 1 are selected at random and copied into the offspring at exactly the same location as they appear in the parent gene. Thus, both order and position of the genes in parent 1 are preserved.
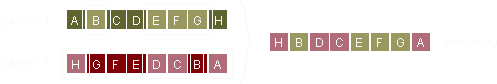
The genes that aren't selected from parent 1 are copied from parent 2 to fill out the empty spaces in the offspring. The order of the genes in parent 2 is preserved but the exact locations are, for obvious reasons, not.
Fixed Position Sequential
One at random selected part (consisting of consecutive genes) of the parent 1 chromosome is copied into the offspring at exactly the same location. The order and location is preserved.
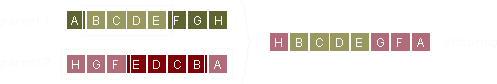
Just as in 'Fixed Position' described above, the parent 2 chromosome is used to fill out the empty locations in the offspring. Order but not location is preserved.
Mutation
One at random selected part (consisting of consecutive genes) of the chromosome is permuted. In practice, this is done by switching location of the genes within the selected part.

Fitness Evaluation
The fitness is calculated in two steps. First, all chromosomes in the population is evaluated. Then, one of the three fitness algorithms are applied.
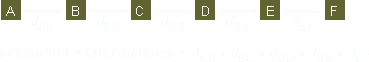
| Chromosome no: | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
| Evaluation value: | 125 | 113 | 092 | 037 | 022 | 018 | 009 | 008 | 005 | 003 |
| Fitness: | 125 | 113 | 092 | 037 | 022 | 018 | 009 | 008 | 005 | 003 |
Windowing
This algorithm translates the entire population fitness so that the weakest chromosome gets fitness 1, and the other gets a fitness equal to the amount by which they exceed the weakest chromosome evaluation.
| Chromosome no: | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
| Evaluation value: | 125 | 113 | 092 | 037 | 022 | 018 | 009 | 008 | 005 | 003 |
| Fitness (guard = 1): | 123 | 111 | 090 | 035 | 020 | 016 | 007 | 006 | 003 | 001 |
| Fitness (guard = 10): | 123 | 111 | 090 | 035 | 020 | 016 | 010 | 010 | 010 | 010 |
The guarding value can be thought of as a social security algorithm for the weak chromosomes. Every chromosome with a fitness value less than the guarding value is assigned the guarding value.
Linear Normalization
The best chromosome is assigned fitness = 100. All other chromosomes' fitness gets decreasing values of the best fitness value. The decrement factor is set to be [x] percent (assigned in the "Parameter" form) of the best fitness which usually results in a slow but stable population fitness convergence.
| Chromosome no: | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 |
| Evaluation value: | 125 | 113 | 092 | 037 | 022 | 018 | 009 | 008 | 005 | 003 |
| Fitness (decr. = 10%): | 100 | 090 | 080 | 070 | 060 | 050 | 040 | 030 | 020 | 010 |
Population Deletion
In this system, two different kinds are implemented:
The "Elitism Value" parameter is used to assure that the best [x] chromosomes will survive the generational replacement.
Steady Delete
In this algorithm, every new offspring is replacing the worst chromosome in the population. Chromosomes that are members of the same generation can be eachother's parents! This is, of course, not biological viable but can have good effect in speeding up the population fitness convergence during evolution.
The program consists of a main part and three classes: Gene, Chromo and Pop.
Gene
This class defines the 'atomic' parts of the GA.
class Gene
{
public:
Gene(char *_name, coord_t _x, coord_t _y,
unsigned int _index, unsigned int _distVecLength);
~Gene();
// Calculates Eucledian distance between this gene and _gene and returns the value.
distance_t calculateDistanceTo(Gene& _gene);
// Returns Eucledian distance between this gene and _gene.
distance_t distanceTo(Gene& _gene);
// Returns this gene's index in original chromosome pool.
unsigned int getIndex() {return index;}
// Returns this gene's name.
char *getName() {return name;}
// Prints gene information to stream.
friend ostream& operator<<(ostream& outStr, Gene& gene);
private:
distance_t *distance; // distance vector
unsigned int distVecLength; // length of distance vector
char *name; // name of gene
coord_t x,y; // gene's coordinates
unsigned int index; // gene's index
};
A Gene consists of a name, x- and y-coordinates, a distance vector describing the distances to other genes (only calculated once at every program execution) and a gene index.
Possible operations on a Gene can be seen in the class declaration above.
Chromo
This class defines what a chromosome looks like.
class Chromo
{
public:
Chromo(unsigned int _chromoLength, Gene *_genePool[]);
~Chromo();
// Copy constructor.
Chromo& Chromo::operator=(Chromo& original);
// Chromosome initialization.
void init();
// Calculates (if necessary) and returns chromosome evaluation.
eval_t eval();
// Sets chromosome's modified evaluation (fitness).
eval_t setModEval(eval_t _modEval) {return (myModEval = _modEval);}
// Returns chromosome's modified evaluation (fitness).
eval_t getModEval() {return myModEval;}
// Returns chromosome's evaluation (fitness).
eval_t getEval() {return myEval;}
// Mates this chromosome with partner resulting in two new offsprings, child1 and child2.
void mate(float _crossover, cross_t crossoverType, float _mutation, Chromo& partner,
Chromo& child1, Chromo& child2);
// Returns chromosome length.
unsigned int getLength() {return chromoLength;}
// Prints chromosome information to stream.
friend ostream& operator<<(ostream& outStr, Chromo& chromo);
private:
unsigned int chromoLength; // chromosome length
eval_t myEval; // chromosome evaluation
eval_t myModEval; // chromosome modified evaluation (fitness)
Gene **genePool; // pointer to original gene pool vector
Gene **myGenePool; // private gene pool vector
// Keeping track of used genes during crossover.
int Chromo::isUsed(Gene *gene, Gene *usedGene[], int noOfUsedGenes);
// Performs crossover using parent1 and parent2 as templates.
// This chromosome is assigned the result.
void crossover(cross_t crossoverType, Chromo& parent1, Chromo& parent2);
// Performs mutation on this chromosome.
void mutate();
// Modified random generator.
unsigned int myRand();
};
A Chromo consists mainly of a Gene vector and two evaluation values (one original and one modified by one of the fitness algorithms).
Possible operations, including crossover and mutation, can be seen in the class declaration above.
Pop
This class defines the population.
class Pop
{
public:
Pop(int _popSize, unsigned int _chromoLength, Gene *_genePool[]);
~Pop();
// Population initialization.
void init();
// Calculates (if necessary) fitness for each chromosome in the
// population and returns the sum of the fitness values.
eval_long_t fitness(fitness_t fitnessType, eval_t fitnessParam);
// Creates a new population using this population as a parent pool
// and returns a pointer to the new population.
// If deletionType == deleteAll, the user of this class has to
// delete this population (the new one is really a _new_ one).
// If deletionType == steadyDelete, the old population is altered
// and thus, requires no further action by the user.
Pop *newPop(fitness_t fitnessType, eval_t fitnessParam,
float crossover, cross_t crossoverType, float mutation,
delete_t deletionType, int elitismValue);
// Returns the fitness of the best cromosome in the population.
eval_t bestFitness() {return myChromoPool[0]->getEval();}
// Returns the modified fitness of the best cromosome in the population.
eval_t bestFitnessMod() {return myChromoPool[0]->getModEval();}
// Returns the average chromosome fitness of the population.
eval_t fitnessAverage();
// Returns the average modified chromosome fitness of the population.
eval_t fitnessAverageMod() {return (eval / popSize);}
// Returns the best chromosome in the population.
Chromo& bestChromo() {return *myChromoPool[0];}
Chromo **myChromoPool; // Chromosome vector
// (has to be public for some reason I can't remember at the time beeing)
// Prints population information to stream.
friend ostream& operator<<(ostream& outStr, Pop& pop);
private:
Gene **genePool; // pointer to original gene pool vector
unsigned int chromoLength; // length of chromosomes in the population
eval_long_t eval; // sum of all chromosome evaluation values
int popSize; // number of chromosomes in the population
// Returns a suitable parent using Roulette Wheel Selection.
Chromo& getParent();
// Sorts chromosome pool in fitness order.
void quickSort(int start, int finish);
};
A Pop mainly consists of a Chromo vector.
Possible operations can be seen in the class declaration above.
Main
The major object for this part is to assign the GA the user parameters and to execute the GA. The GA execution loop looks like this:
//******************* Main Evolution Loop **********************
Pop *myPop = new Pop(popSize, chromoLength, genePool);
Pop *myNewPop;
int k;
cout << "Data:" << endl;
for (k = 0; k < generations; k++)
{
myPop->fitness(fitnessType, fitnessParam);
cout << k << " "
<< myPop->fitnessAverage() << " "
<< myPop->bestFitness() << endl;
myNewPop = myPop->newPop(fitnessType, fitnessParam, crossover,
crossoverType, mutation, deletionType,
elitismValue);
if (deletionType == deleteAll)
{
delete myPop;
myPop = myNewPop;
}
}